35.0 - Create/delete Contacts
Contacts can be formed between two pieces of different bodies, and the pieces involved in a contact are called the contact ends. Prior to the first cycle in a series of cycles, all contacts are created between pieces (during the initialization of data structures as shown here). This is necessary because without contacts, one cannot calculate the effective stiffness of the system, meaning that a stable timestep cannot be computed.
Contact updating is triggered for pieces that were deemed dirty by their cumulative motion exceeding their tolerance extent criteria. Such pieces have been remapped in the cell space during cycle point 30.0 (see here for details). Contacts are created when the piece conformal surfaces overlap and are deleted when the conformal surfaces no longer overlap. Contact updating only occurs for dirty pieces. Note that one can ensure contacts are never deleted by setting the state of the contact to always active with the contact.activate FISH function.
With the timestep determination mode set to either automatic or scaling (see the model mechanical timestep command), the timestep is limited so that any piece may, at most, translate \(\varepsilon\) from its last position. Given the enforcement of this constraint, one can guarantee that contacts are created prior to the cycle where forces/moments exist. This is pictorially shown below. Suppose that two identical balls (identical geometries and \(\varepsilon\) values) exist in such a configuration that their extents lie infinitesimally close (by a distance of \(\vartheta\)) at the end of cycle \(N\). Additionally, suppose that both balls have velocities \(\left | \mathbf{v} \right | = (\varepsilon-\vartheta) / \Delta t\) directed toward one another, where \(\Delta t\) is the timestep during cycles \(N+1\) and \(N+2\). At the end of cycle \(N+1\), the extents remain unchanged because neither ball has displaced \(\varepsilon\) from its cycle \(N\) position. The ball locations are updated during cycle \(N+2\) such that the balls are \(\overline{r}_i + \vartheta\) from one another. Subsequently, the piece extents are updated and the contact is created prior to the cycle when forces/moments could potentially develop.
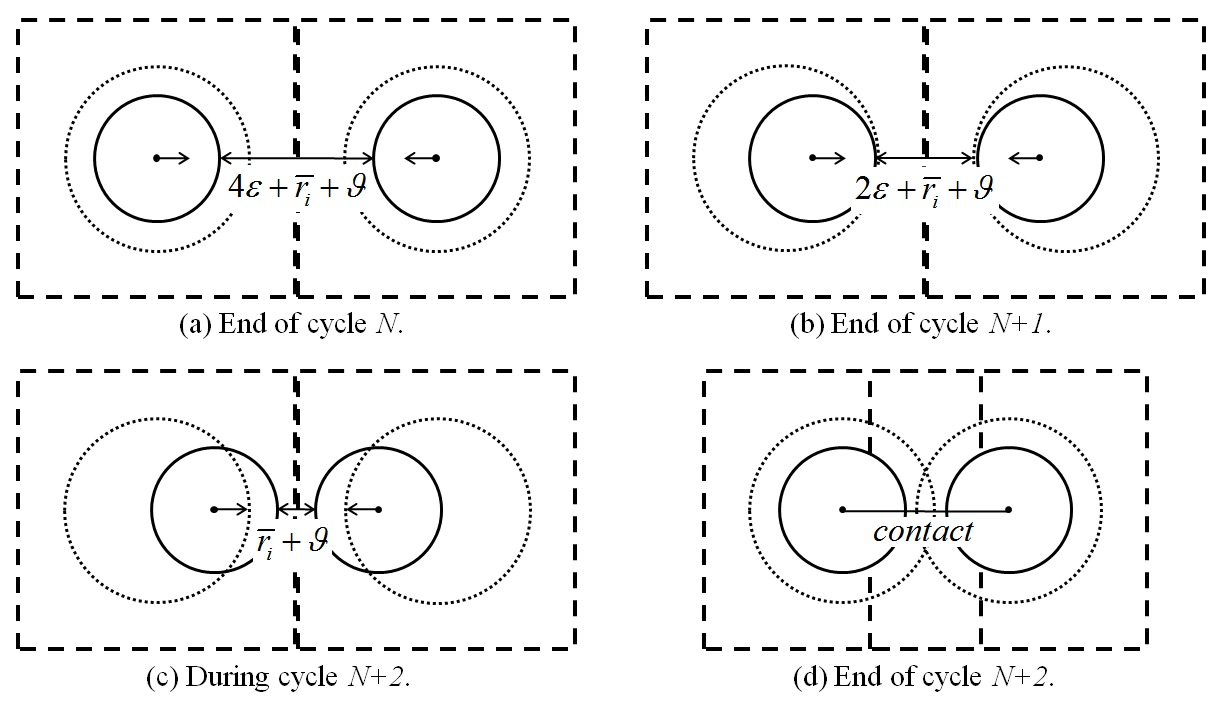
Figure 1: Demonstration of the translation-limiting logic for identical pieces (solid lines) moving toward one another with fixed velocities. The cell extents (dashed lines) are updated when the piece displacements exceed \(\varepsilon\) (small-dashed lines).
Upon creation, a contact model is assigned to each contact based on the current configuration of the Contact Model Assignment Table (CMAT). As described in the “Model Components” section, the contact geometry is delineated upon creation.
| Was this helpful? ... | PFC © 2021, Itasca | Updated: Feb 25, 2024 |
